Data tables are becoming an increasingly popular way of working with data sets in R. The syntax can become rather complex, but the framework is much faster and more flexible than other methods. The basic structure of a data table, though, if fairly intuitive–as it corresponds quite nicely with SQL queries in relational database management systems. Here’s the basic structure of a data table:
DT[i, j, by =],
where ‘i’ = WHERE, ‘j’ = SELECT, and by = GROUP BY in SQL
The easiest way to explain how the data table framework in R is like SQL, though, is to offer an example. So, that’s what we’re going to do. Let’s start by pulling a data set in R…
### bring data into R with data.table
install.packages("data.table")
library(data.table)
profs <- fread("https://vincentarelbundock.github.io/Rdatasets/csv/car/Salaries.csv")
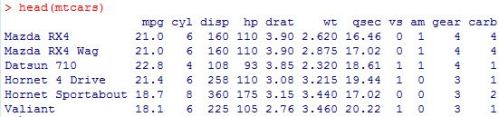
head(profs)
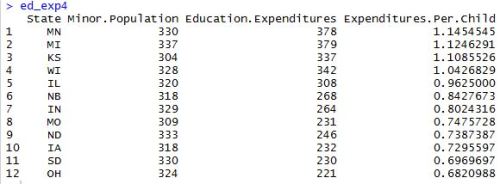
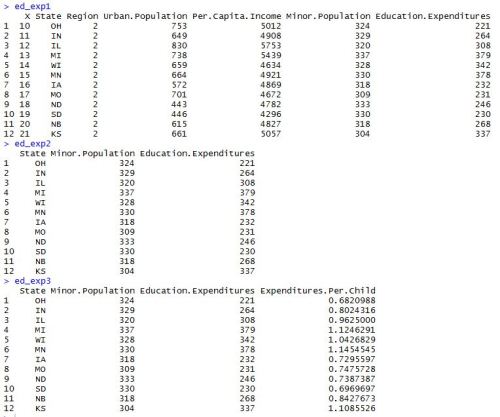
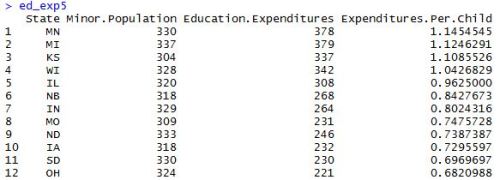
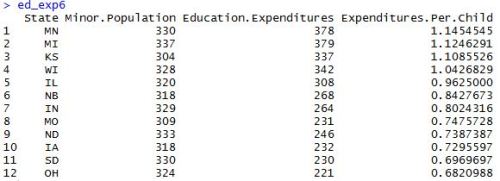
After installing and loading the data.table package, we’ll be able to use the fread() function to import a data set as a data table. When we run the head() function on the profs data table, we can see what we’re looking at. We’ve got a collection of college profressors’ salaries based on a variety of factors.

Now, suppose we want to call the same information in MySQL. Here’s the code for that…
### select the first 6 rows of the profs table select * from profs limit 6;
And here’s the results of the query shown in the MySQL Workbench.
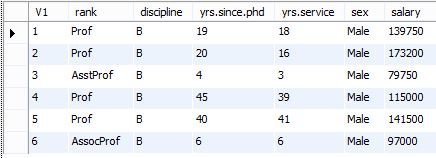
Pretty simple, right?
Now, let’s suppose we have a specific question we want to answer. What if we want to know whether or not there is a difference between salaries for males and females when they become full professors. How do we use a data table to do this in R. Here’s what the code looks like…
### find average full professor salary by gender profs[rank == "Prof", .(avg.salary = mean(salary)), by = sex][order(-avg.salary)] ### step by step with notes comparing to my SQL profs[ ### the [] subsetting of profs corresponds to the from clause in MySQL rank == "Prof", ### selects rows where rank is professor; corresponds to where clause in MySQL .(avg.salary = mean(salary)), ### selects the columns to return; corresponds to select clause in MySQL by = sex ### corresponds to the group by clause in MySQL ][order(-avg.salary)] ### adding a second [] subset with the order function corresponds to the order by clause in MySQL; the '-' in front of the ordered variable corresponds to the desc statement in MySQL
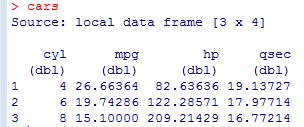
So, as the comments suggest, the three sections within the square brackets of the data table call correspond to what we want to do with the rows, columns, and groupings, respectively. In this scenario, we first select the rows of full professors, then select a column averaging the salaries of full professors, and finally group those full professors by gender. And, here’s what the output looks like.

If we want to write a query to retrieve this same information in MySQL, here’s what the code would look like…
### find average full professor salary by gender select sex, avg(salary) from profs where rank = 'Prof' group by sex order by salary desc; ### step by step with notes comparing to R's data.table select sex, avg(salary) ### selects the columns to return; corresponds to the 'j' section of R's data.table from profs ### selects the table to pull data from; corresponds to the [] subsetting in R's data.table where rank = 'Prof' ### selects the rows to return; corresponds to the 'i' section of R's data.table group by sex ### groups the table by gender; corresponds to the 'by' section of R's data.table order by salary desc; ### orders the table by highest salary to lowest; corresponds to the the order function in R's data.table
And here’s the result in the MySQL Workbench.
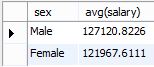
So, to answer our question, male full professors in our data set do indeed make slightly more on average than females who have risen to the same level in their careers.
Go ahead and experiment. What other subsets can you create from this data set using the data.table framework?